This is a presentation by Dominique Estival, Steve Cassidy, Peter Sefton, Denis Burnham and Jared Berghold given at the eResearch Australasia 2013 Conference in Brisbane by Peter Sefton. The presentation is built around a demo of Version 2 of the lab by Steve Cassidy.
Dominique Estival,1 Steve Cassidy2, Peter Sefton1, Denis Burnham1, Jared Berghold3
1 University of Western Sydney, Australia, {d.estival,p.sefton,d.burnham}@uws.edu.au
2 Macquarie University, Sydney, Australia,
[
steve.cassidy@mq.edu.au
](mailto:steve.cassidy@mq.edu.au)
3 Intersect Australia Ltd,
[
jared.berghold@intersect.org.au
](mailto:jared.berghold@intersect.org.au)
##
Administered by MARCS Institute at the University of Western Sydney, the Human Communication Science Virtual Laboratory
HCS vLab
project [
1
] started in December 2012 and is expected to be completed and operational by July 2014. The
HCS VLab
builds upon collaborations set up during the very successful ARC (Australian Research Council)-funded HCS Network project[
2
]; it is the first time such a project has been conducted in Australia and will benefit researchers from myriad disciplines encompassed by Human Communication Science (HCS).
The
HCS vLab
is funded by NeCTAR, a body set up by the Australian Government as part of the Super Science initiative and financed by the Education Investment Fund.
 University of Western Sydney, Macquarie University, the Australian National University, University of Canberra, Flinders University, University of Melbourne, University of Sydney, University of Tasmania, University of New South Wales, University of Western Australia, RMIT, University of New England, LaTrobe University, NICTA (National ICT Australia, ASSTA (Australasian Speech Science and Technology Association),
University of Western Sydney, Macquarie University, the Australian National University, University of Canberra, Flinders University, University of Melbourne, University of Sydney, University of Tasmania, University of New South Wales, University of Western Australia, RMIT, University of New England, LaTrobe University, NICTA (National ICT Australia, ASSTA (Australasian Speech Science and Technology Association),
Link data to tools
Provide a collaboration environment
Improve reusability and replicability of results
Research conducted in isolation entails inefficient repetition of analysis of local data sets. HCS research in Australia, and successful real-life applications, requires going beyond the isolated desk-PC-lab-university-bound model of research into a new research environment. Such an environment will eradicate the waste involved in repeated unshared analyses; ignite the research spark that affords the serendipity of new tool-corpus combinations; and dramatically improve scientific replicability by moving corpora and tools and the analyses conducted with these into an easy access, shared, in-the-cloud, public, replicable environment.
The aims are to:
facilitate access by the Australian and international HCS communities to data and analysis tools;
afford new tool–corpus combinations and new emergent research output –projects, grant funding, doctoral theses, and publications;
allow analysis and annotation results to be stored and shared, thus promoting collaboration between institutions and disciplines;
improve scientific replicability by moving local and idiosyncratic desktop-based tools and data to an accessible, in-the-cloud, environment that standardises, defines, and captures procedures and data output so that research publications can be supported by re-runnable re-usable data and coded procedure (see, e.g., www.myexperiment.org/). The HCS vLab is designed to make use of national infrastructure – including data storage, discovery and research computing services. It incorporates existing eResearch tools, adapted to work on shared infrastructure, with a data-discovery interface to connect researchers with data sets, orchestrated by a workflow engine with both web and command line interfaces to allow use by technical and non-technical researchers, via a Web interface.
 In this presentation we will walk through a demonstration of HCS vLab version 2.0 by Steve Cassidy, showing the architecture of the lab as he uses it. There will not be time in the presentation at eResearch Australasia to watch the full demo.
In this presentation we will walk through a demonstration of HCS vLab version 2.0 by Steve Cassidy, showing the architecture of the lab as he uses it. There will not be time in the presentation at eResearch Australasia to watch the full demo.
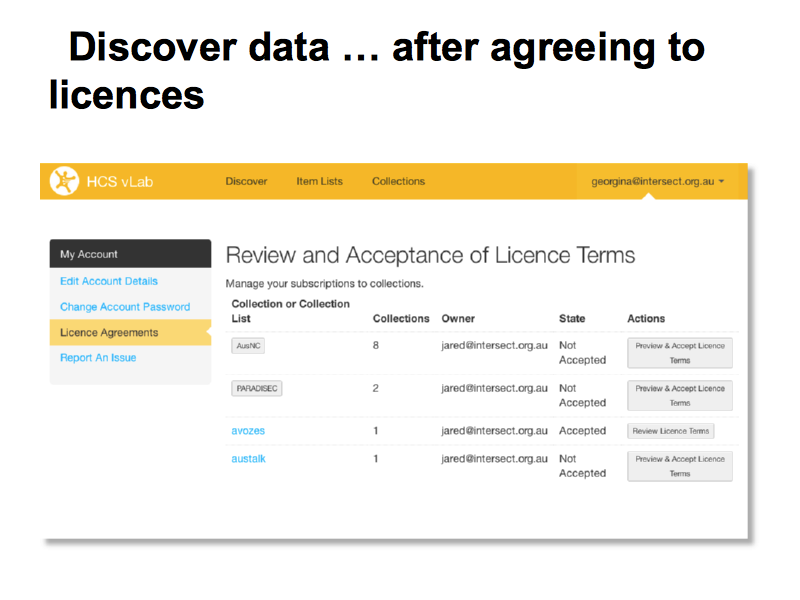 Most of the data sets require users to agree to a click-through license because they involve human participants and because of the arrangements under which they are collected. Some of the licenses require data-owner approval. The entire lab is built around an Application Programming Interface (API) which respects the licenses and gives researchers access to data sets for which they are licensed, through the web, the workflow tool and via programmatic use of the API.
Most of the data sets require users to agree to a click-through license because they involve human participants and because of the arrangements under which they are collected. Some of the licenses require data-owner approval. The entire lab is built around an Application Programming Interface (API) which respects the licenses and gives researchers access to data sets for which they are licensed, through the web, the workflow tool and via programmatic use of the API.
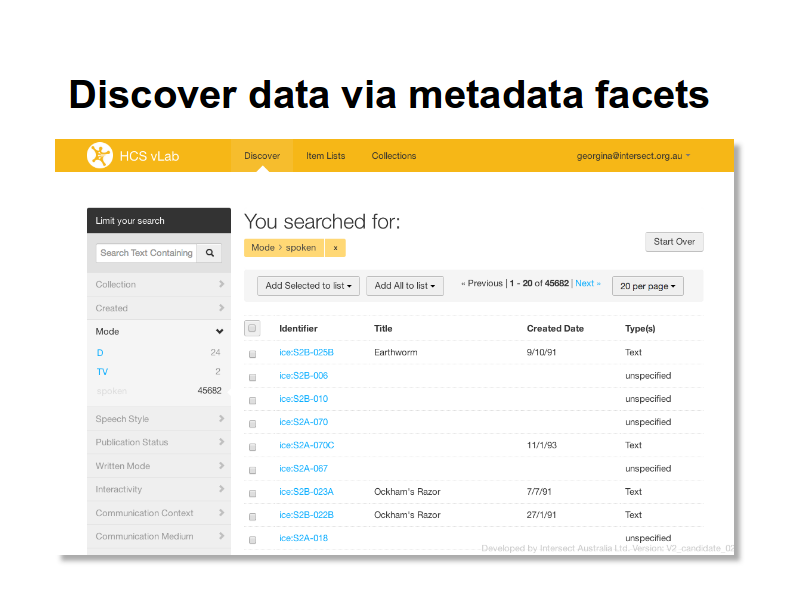 The discovery interface is build on a an Apache Solr index, using the Blacklight project’s user interface framework. This is just the first of many indexes that are planned for the lab.
The discovery interface is build on a an Apache Solr index, using the Blacklight project’s user interface framework. This is just the first of many indexes that are planned for the lab.
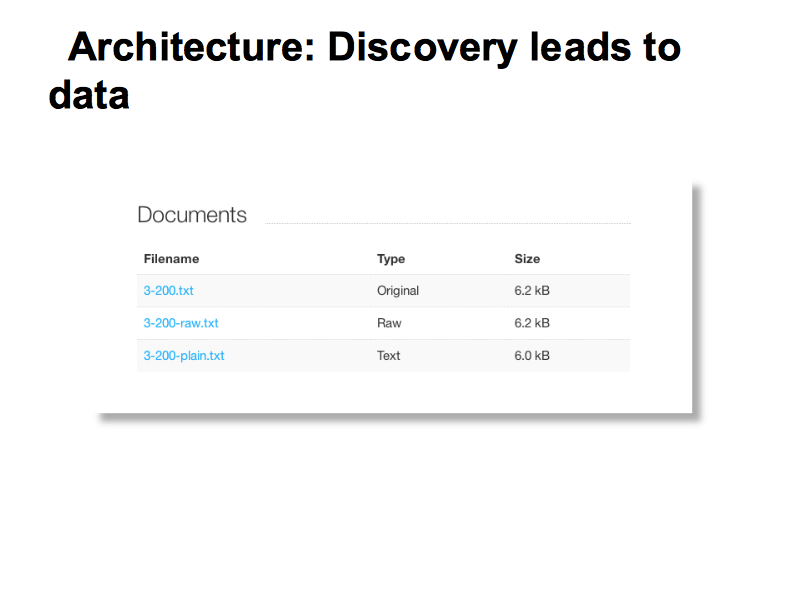 The lab has a data-storage component which can look after large amounts of heterogeneous data.
The lab has a data-storage component which can look after large amounts of heterogeneous data.
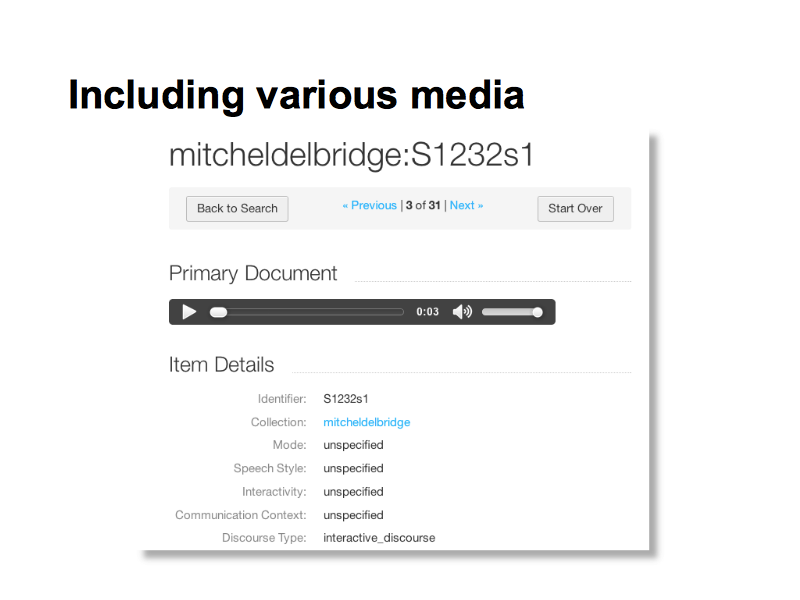 This is an example of speech data from the Mitchell and Delbridge corpus (data set) collected in the 1960s.
This is an example of speech data from the Mitchell and Delbridge corpus (data set) collected in the 1960s.
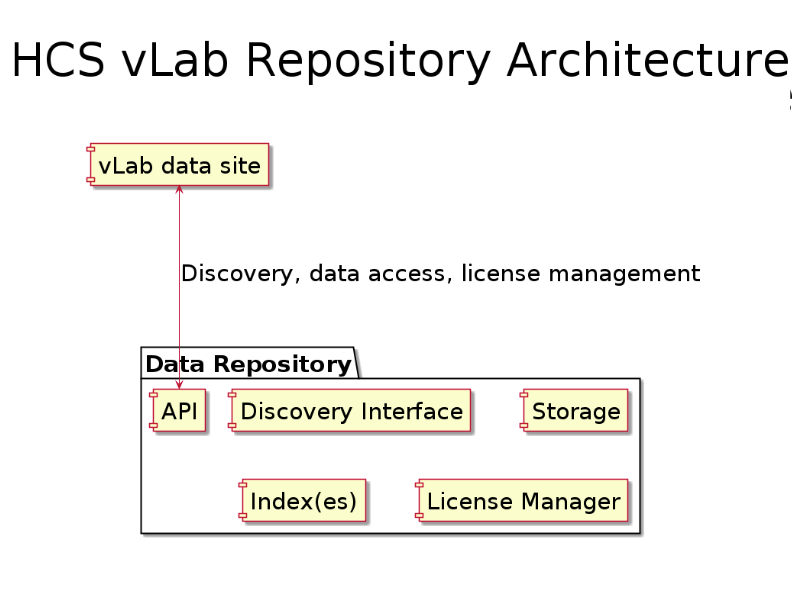 The architecture so far: the HCS vLab website is using a data repository with a series of services:
The architecture so far: the HCS vLab website is using a data repository with a series of services:
* Data storage
* An index of metadata (this one is Apache Solr, more will be added, eg Indri and an RDF triple-store)
* A license manager, integrated with the indexing and storage
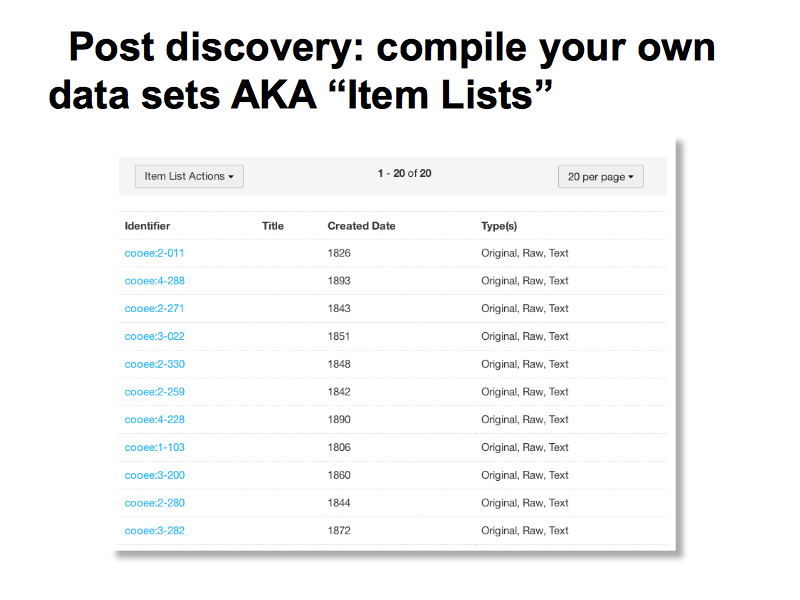 Researchers can create their own data sets from items in the repository. These are know as item lists and these are the key to re-doable research workflows as they allow the same stable data set to be run through multiple processes. Item lists are available via the web interface and via the API.
Researchers can create their own data sets from items in the repository. These are know as item lists and these are the key to re-doable research workflows as they allow the same stable data set to be run through multiple processes. Item lists are available via the web interface and via the API.
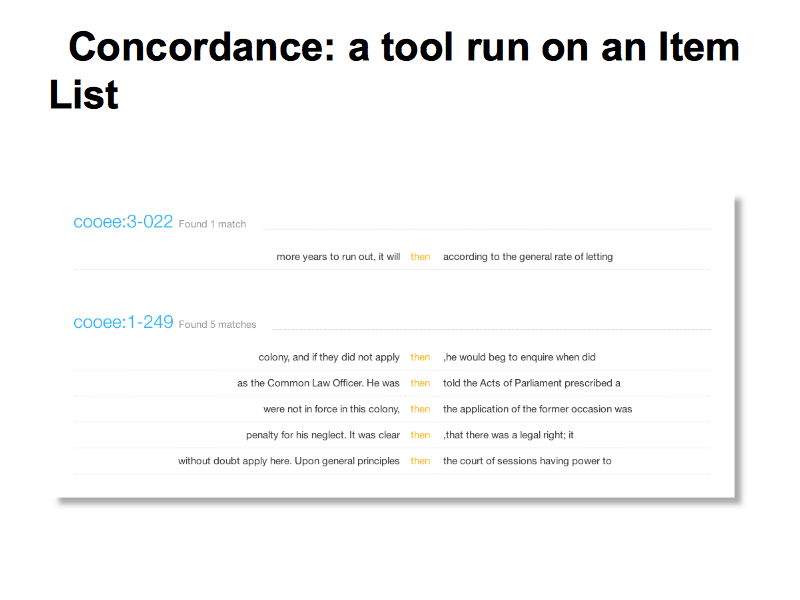 Steve’s demo shows some example of simple, broadly useful text analysis tools that are built in to the HCS vLab data site.
Steve’s demo shows some example of simple, broadly useful text analysis tools that are built in to the HCS vLab data site.
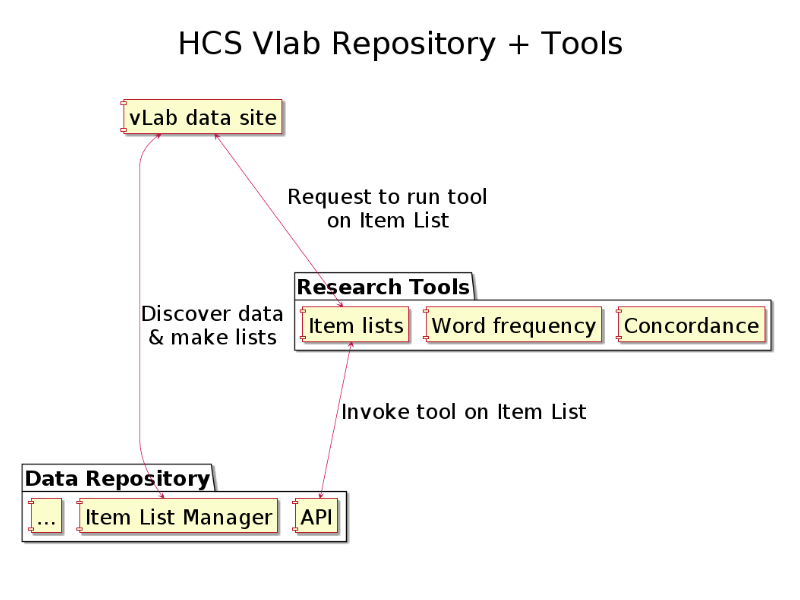 This diagram shows how item lists are the the link between tools and data.
This diagram shows how item lists are the the link between tools and data.
 All the data and discovery services available in the HCS vLab are available via the API.
All the data and discovery services available in the HCS vLab are available via the API.
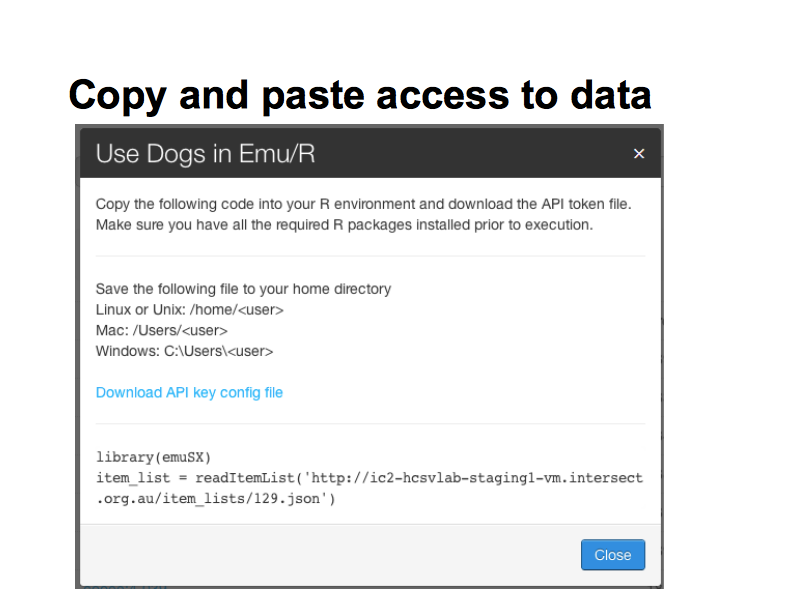 Here we see item list “Dogs” selected – the user is presented with a fragment of R code which will load it into an R data structure.
Here we see item list “Dogs” selected – the user is presented with a fragment of R code which will load it into an R data structure.
API supports rich queries on the repository index from anywhere.
creating and consuming stable item-lists
Fetching data
Secure access based on use license
COMING SOON data deposit and annotation support
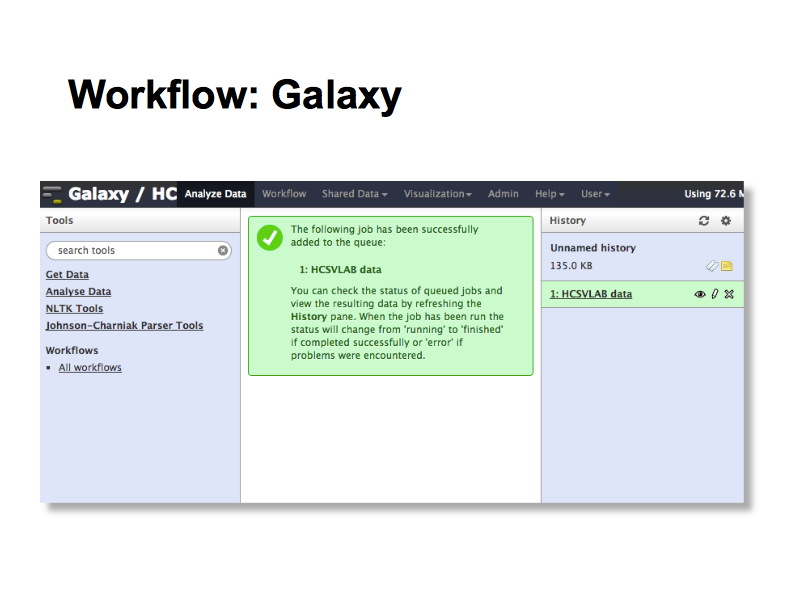 Like other NeCTAR virtual labs, HCS vLab uses the Galaxy workflow engine to allow researchers to construct research workflows that can be run over different data sets, and share them with others. In phase two of the project we will work on reusablity and reproducibility with research teams.
Like other NeCTAR virtual labs, HCS vLab uses the Galaxy workflow engine to allow researchers to construct research workflows that can be run over different data sets, and share them with others. In phase two of the project we will work on reusablity and reproducibility with research teams.
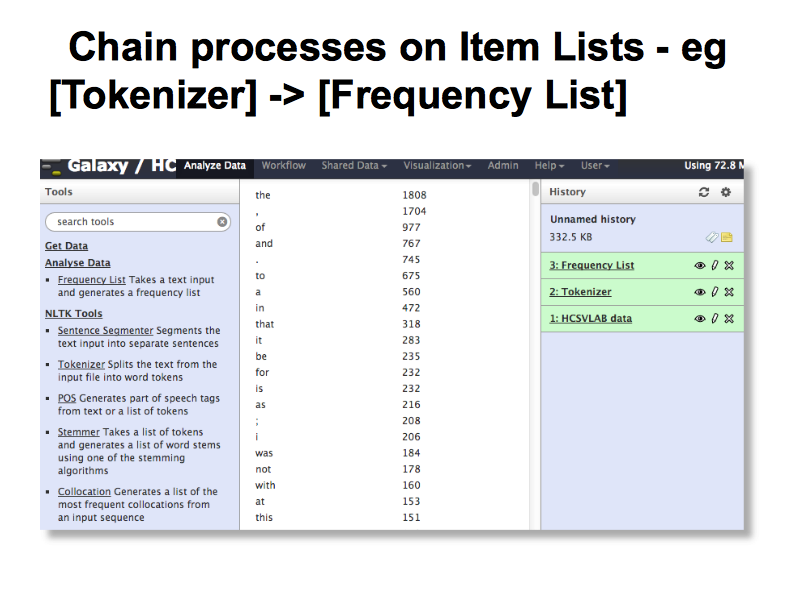 This simple demo show how an Item List is used in Galaxy, where multiple operations can be performed on data with each step feeding its result into the next one. This allows for more flexibility than the sinple ‘canned’ tools built into the HCS vLab data web site.
This simple demo show how an Item List is used in Galaxy, where multiple operations can be performed on data with each step feeding its result into the next one. This allows for more flexibility than the sinple ‘canned’ tools built into the HCS vLab data web site.
A similar approach will be used for audio and video data allowing researchers to create item lists, set some parameters and run analytical processes, resulting in new data sets or graphical plots.
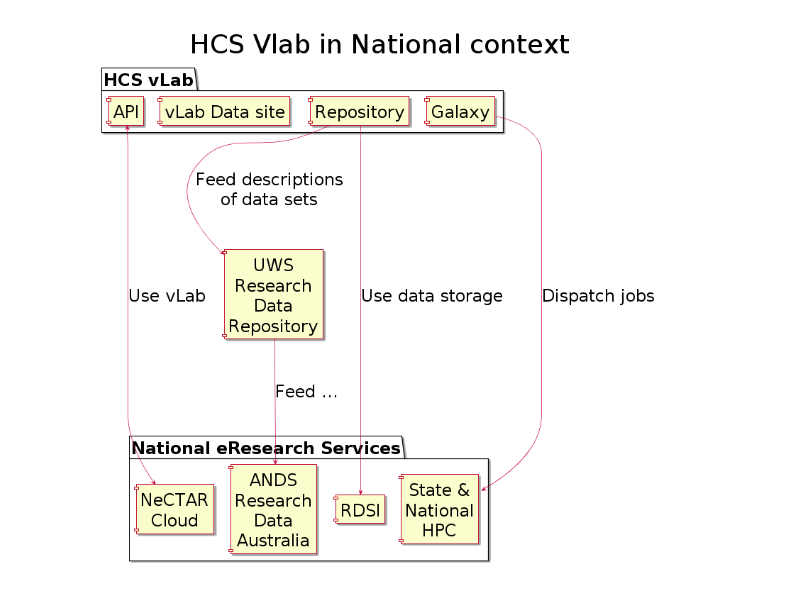
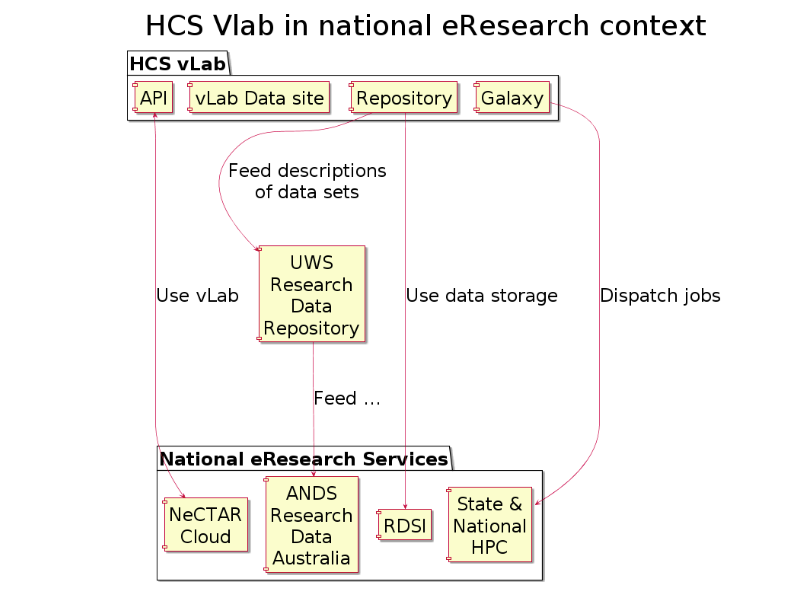 No vLab is an island: the virtual laboratory is plugged-in to the national eResearch service-grid.
No vLab is an island: the virtual laboratory is plugged-in to the national eResearch service-grid.
 This project uses a number of open source components to scaffold the lab including
This project uses a number of open source components to scaffold the lab including
Hydra which is Ruby on Rails framework wrapping:
Apache Solr and blacklight
The Duraspace Fedora Commons repository
Apache Solr, via the Blacklight project
Workflow courtesy of Galaxy “Data intensive biology
for everyone.” 🙂
 ## “The HCS vLab is easily one of the best interfaces I’ve come across in Australian research projects and eResearch applications. The Bootstrap framework is a great development platform and the workflows and interfaces of the HCS vLab have been integrated well to form a beautifully clean and usable application.”
## “The HCS vLab is easily one of the best interfaces I’ve come across in Australian research projects and eResearch applications. The Bootstrap framework is a great development platform and the workflows and interfaces of the HCS vLab have been integrated well to form a beautifully clean and usable application.”
AeRO is “Australian eResearch Organisations”
 ## “I really liked using the system and the instructions were very easy to use and the system easy to navigate. […] This platform would be very useful for my research.”
## “I really liked using the system and the instructions were very easy to use and the system easy to navigate. […] This platform would be very useful for my research.”
–Tester
“It seemed pretty easy to use after I got used to the two platforms. I would certainly like to use it in my future research!”
“It is looking very good. Lots of possible uses and a nice interface.”
“The platform is easy to use and has the great potential to help with Linguistic research and wide applications in other areas…”
“The system seemed to be quite user-friendly. As first I was relying on the manual, however when the manual became more streamlined with less details, the system was still easy to follow.”
“This is a powerful tool and I think it is pretty good.”
“I really liked using the system and the instructions were very easy to use and the system easy to navigate. […] This platform would be very useful for my research.”
“I think it’s quite easy to use. […] Generally the platform is very clearly organised, and user-friendly.”
“The platform overall is very good.”
“Very nice platform with great user interface!”
“A very promising and impressive setup so far!”
“I’m impressed with the platform – it’s smooth and the interface is very intuitive.”
“I think it’s quite easy to use. […] Generally the platform is very clearly organised, and user-friendly.”
“The platform overall is very good.”
“Very nice platform with great user interface!”

This work by Dominique Estival, Steve Cassidy, Peter Sefton, Denis Burnham and Jared Berghold is licensed under a Creative Commons Attribution 3.0 Unported License.